RStudio Server
The RStudio Server interactive app launches a private RStudio IDE session that runs on an ACE HPC compute node and streams to your browser. You get the full RStudio experience — console, script editor, plots pane, environment inspector — without installing anything locally.
Launching RStudio Server
- In the navigation bar, click Interactive Apps
- Select RStudio Server
- A launch form opens — fill in the resource fields (described below)
- Click Launch
- Wait for the session to start — this usually takes 30 seconds to a few minutes depending on queue wait time
- When the session is ready, click the Connect to RStudio Server button
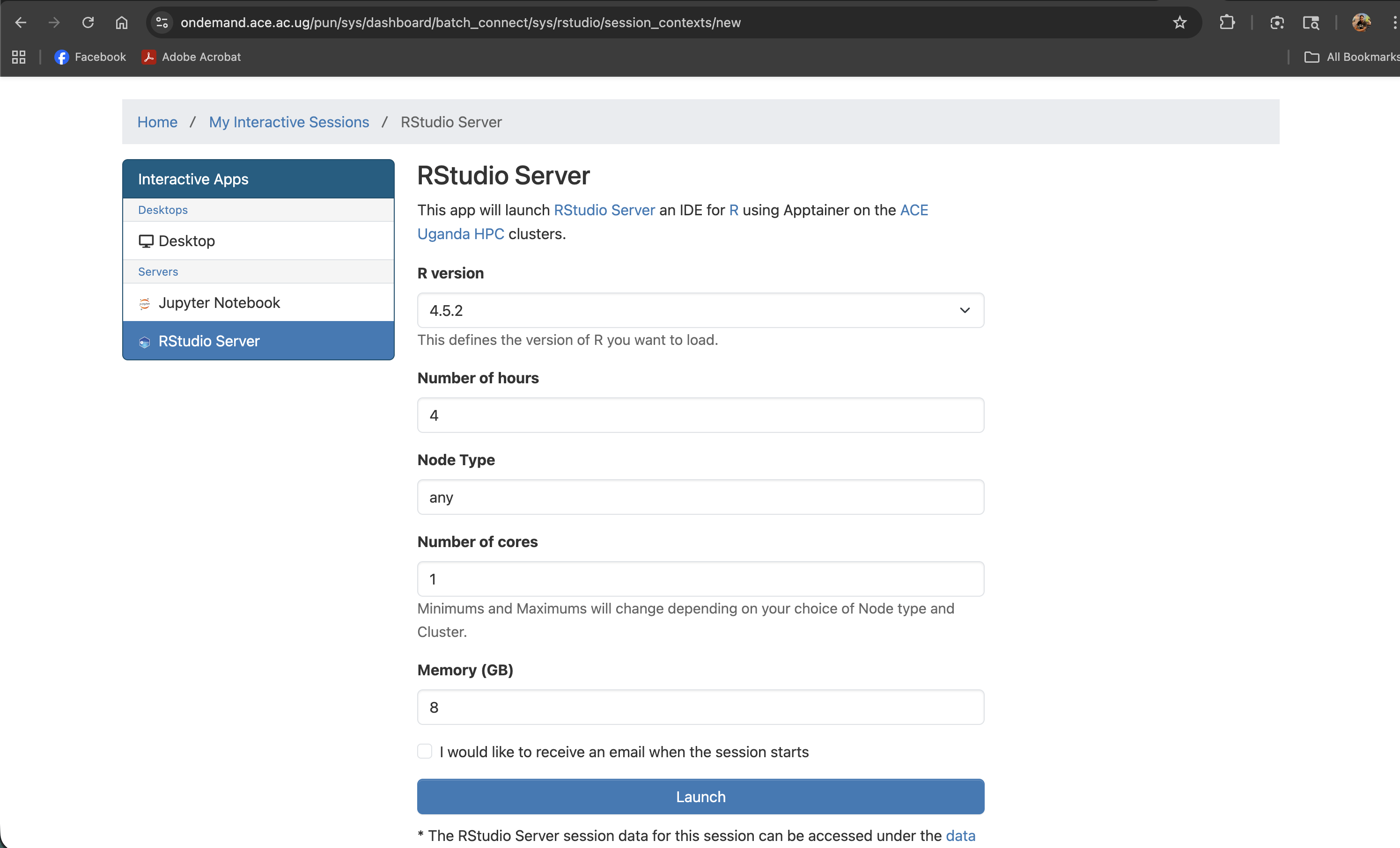
Screenshot note: A screenshot of the RStudio Server launch form with all fields visible before submitting.
Screenshot note: A screenshot of the "Session starting" / queued status card on the dashboard.
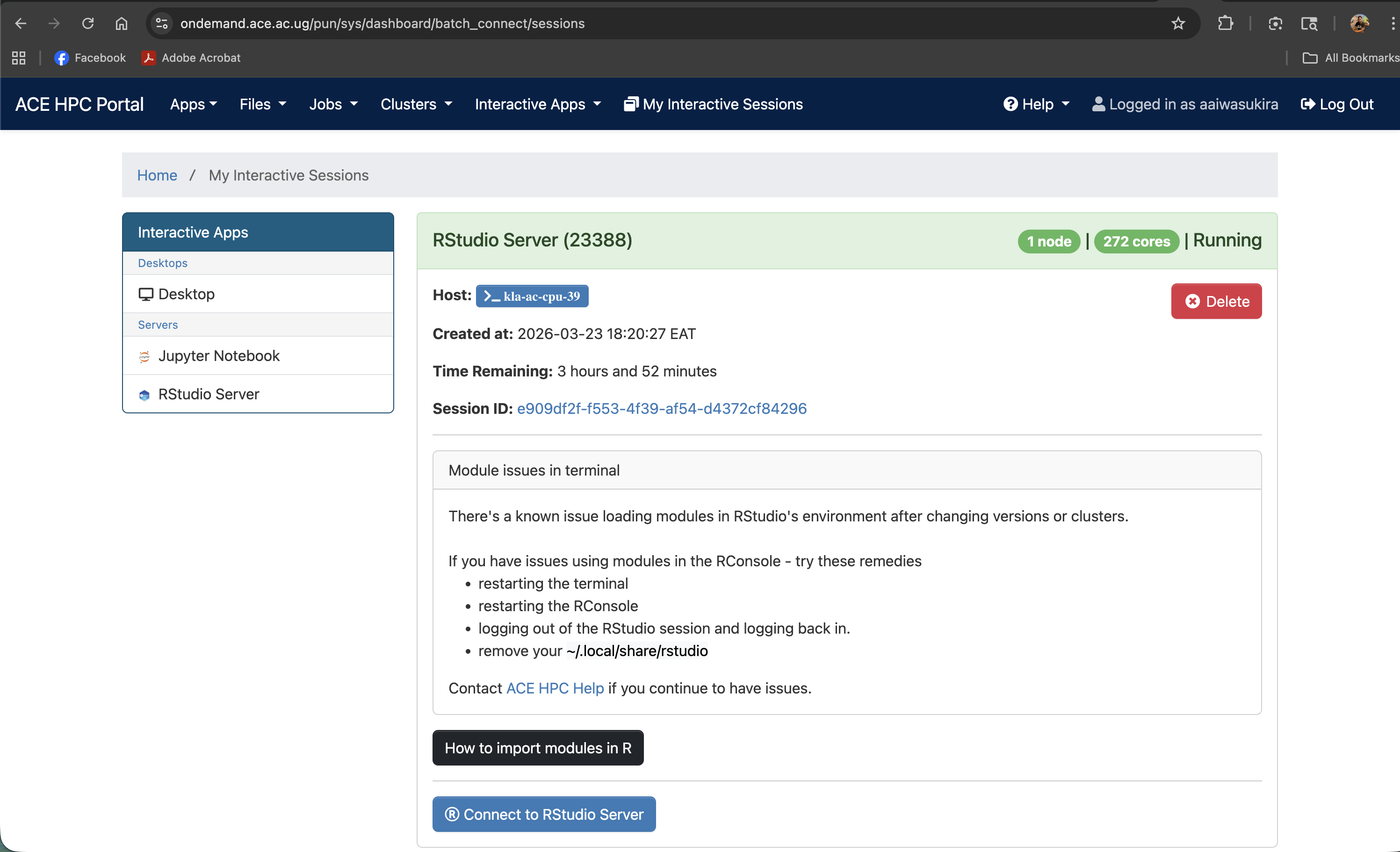
Screenshot note: A screenshot of the "Connect to RStudio Server" button when the session is ready.
Understanding the Launch Form
Every field in the launch form maps directly to a SLURM resource request. Setting these thoughtfully ensures your job starts quickly and has the resources it needs.
R version
R version: [ 4.5.2 ▼ ]
A dropdown listing the R versions installed on ACE HPC. Select the version your project requires. If a version you need is not listed, contact the support team.
Number of hours
Number of hours: 4
The wall-time limit for your session. When this time expires RStudio is shut down automatically and any unsaved work in memory is lost. Your files on disk are always safe.
| Analysis type | Suggested hours |
|---|---|
| Exploratory data analysis | 2–4 |
| DESeq2 / edgeR pipeline | 4–8 |
| Large genome-wide analyses | 8–24 |
You can always delete the session early to free resources, but you cannot extend it once it has started.
Node Type
Node Type: any
The type of compute node to run your session on. Leave this as any unless you have a specific reason to target a particular node type (e.g., a high-memory node). The minimums and maximums for cores and memory vary by node type — setting any lets the scheduler place your job on the first available node that satisfies your resource request.
Number of cores
Number of cores: 1
CPU cores allocated to your RStudio session. Most R code is single-threaded by default, but packages like BiocParallel, doParallel, furrr, and data.table can use multiple cores.
| Scenario | Recommended cores |
|---|---|
| Standard analysis / plotting | 1–2 |
Parallel computing with BiocParallel or foreach | 4–8 |
The available range changes depending on the Node Type selected above. Requesting more cores than your analysis uses wastes resources and can delay your job's start. Set cores equal to what you will actually pass to BiocParallel::MulticoreParam(workers = N) or similar.
Memory (GB)
Memory (GB): 8
RAM allocated to your session. R loads entire datasets into memory, so this is often the most important field to set correctly.
| Dataset / analysis type | Suggested memory |
|---|---|
| Small datasets (< 1M rows) | 4–8 GB |
| Bulk RNA-seq (DESeq2, edgeR) | 16–32 GB |
| Single-cell RNA-seq (Seurat, Bioconductor) | 32–64 GB |
| Genome-wide association studies | 64–96 GB |
R typically uses 2–5× the raw file size in memory. A 5 GB count matrix can use 10–25 GB of RAM.
Email notification
[ ] I would like to receive an email when the session starts
Tick this checkbox if you want to receive an email notification when your RStudio session is ready. This is useful when you expect a queue wait — you can do other work and come back when the email arrives.
Using RStudio in the Browser
Once connected, the interface is identical to desktop RStudio:
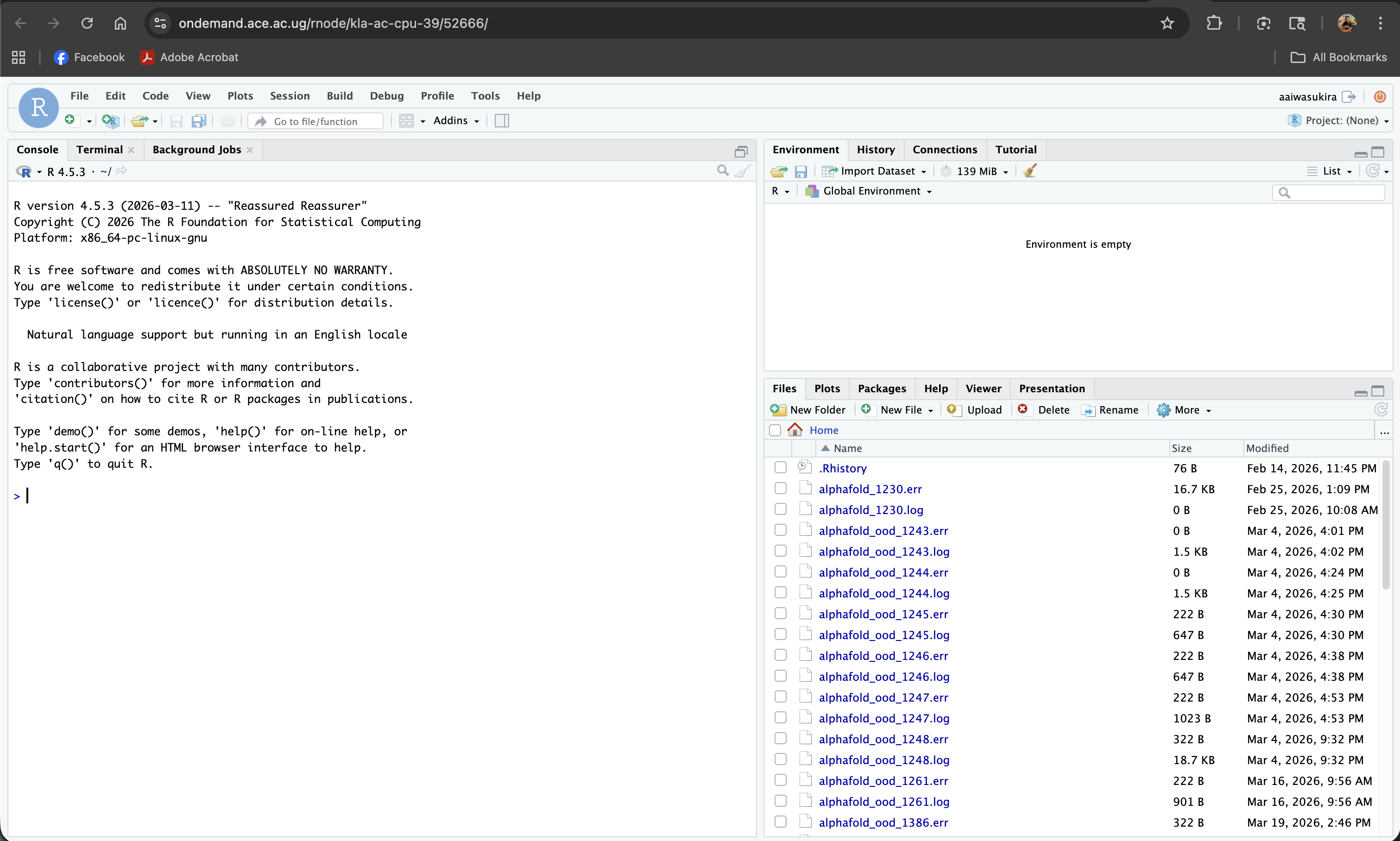
Screenshot note: A screenshot of the full RStudio interface running in the browser after a session connects.
Your working directory
RStudio sessions start with your home directory as the working directory. To confirm:
getwd()
# [1] "/home/username"
For analyses that read large data files, change your working directory to your scratch space:
setwd("/etc/ace-data/home/username/my_project")
Or set it through the Files pane: navigate to the folder, then click More → Set As Working Directory.
Accessing your files
The Files pane (bottom-right) shows the contents of your current working directory on the ACE HPC filesystem. You can navigate it just like a file browser, open scripts by clicking them, and use Upload to transfer small files from your local machine.
For large file transfers, use the OOD File Manager or scp/rsync over SSH.
Installing R packages
Install packages as usual from the R console:
install.packages("ggplot2")
# Bioconductor packages
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("DESeq2")
Packages are installed into your personal library in ~/R/ and persist between sessions. You do not need to reinstall packages every time you launch a new session.
Package installation from CRAN or Bioconductor requires internet access from the compute node. On ACE HPC, compute nodes have outbound internet access for package downloads. If an install fails with a network error, contact support.
Saving your work
- Scripts: save with
Ctrl+S(orCmd+Son macOS) — they are saved directly to the cluster filesystem - RData / RDS files: use
save.image(),save(), orsaveRDS()to write R objects to disk - Plots: use Export in the Plots pane, or
ggsave()/pdf()in code to write to the filesystem
There is no autosave of in-memory R objects. If the session times out or you lose connection without saving, unsaved R objects are lost. Regularly save intermediate results with saveRDS().
Ending Your Session
When you are done:
- Save all open scripts and write any important R objects to disk
- Return to the Open OnDemand dashboard tab in your browser
- Find your session in the My Interactive Sessions panel
- Click Delete to terminate the session and release the compute node
Leaving an idle RStudio session running holds a SLURM allocation, preventing other users from using those cores and memory. Please delete your session as soon as you are finished.